What's New in HumanBase
February 2026
HumanBase Published in Nature Methods:
We're thrilled to share that HumanBase has been published as a Correspondence in Nature Methods! This paper represents years of development and collaboration, and we're grateful to the research community that has made HumanBase a part of their work — over 20,000 researchers across 110 countries in the past year alone.
If HumanBase has contributed to your research, we'd appreciate your citation:
Sealfon, R.S.G., Theesfeld, C.L., Funk, J., Sauerwald, N., Wong, A.K. & Troyanskaya, O.G. HumanBase: an interactive AI platform for human biology. Nat Methods (2026). https://doi.org/10.1038/s41592-025-02994-8
Stay tuned — we have exciting updates in the pipeline, including expanded network coverage and new analysis capabilities coming in the weeks ahead.
August 2025
Updated Downloads Section:
We have redesigned the Downloads page to provide better organization and access to HumanBase resources. The page now includes new bulk download zip files for all top edges, full networks, and gold standards, as well as a Python script for downloading all tissue-specific networks incrementally with error handling. It also features downloadable resources for HumanBase variant prediction tools.
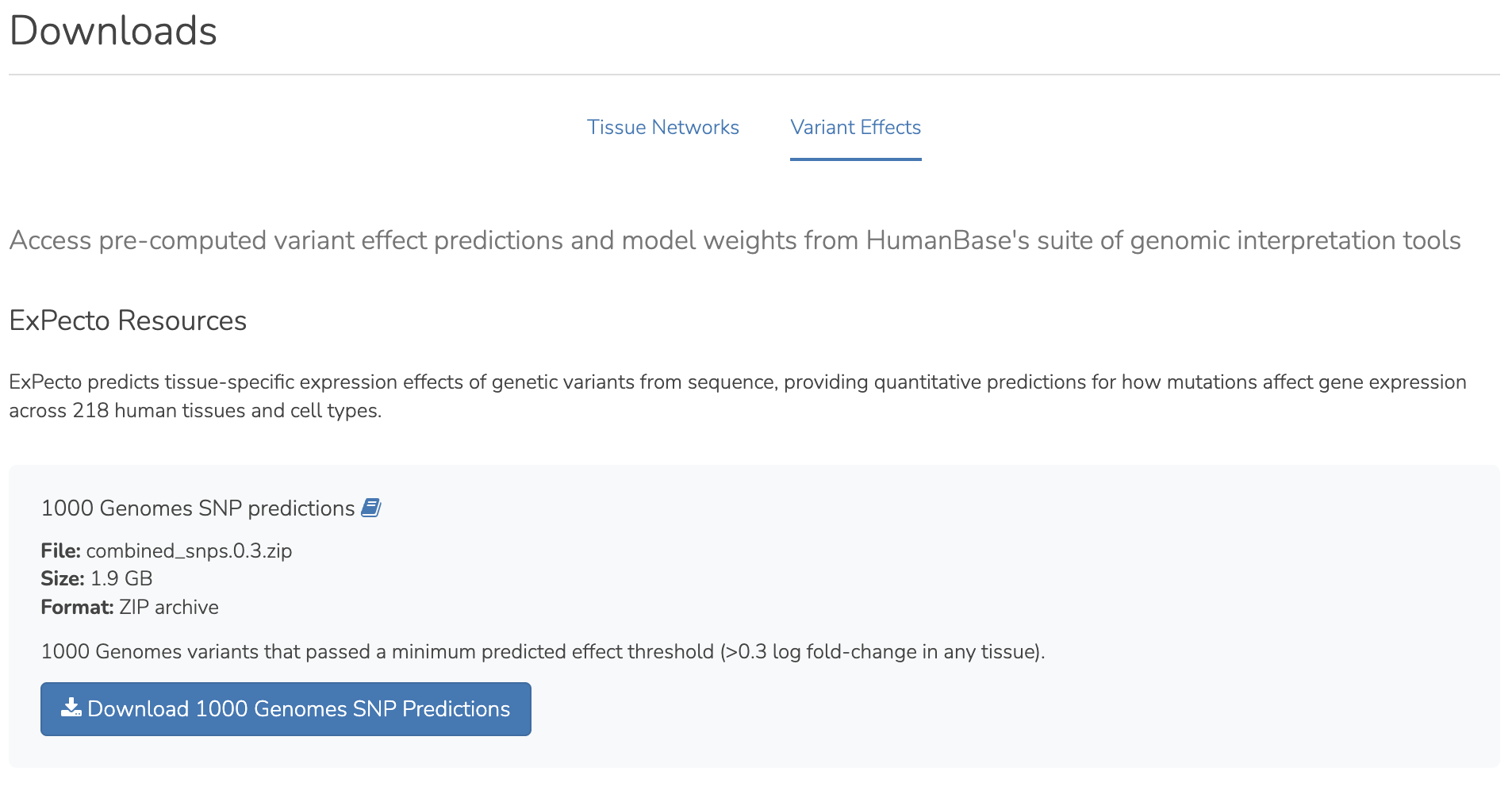
Use Cases:
We have added an extensive collection of use cases/tutorials to the interface demonstrating real research workflows for variant effect prediction and network analysis frameworks.
Enhanced Score Documentation:
We have expanded documentation of score types reported by HumanBase variant effect prediction frameworks and provided tooltip links to relevant score documentation sections from the interface.
October 2024
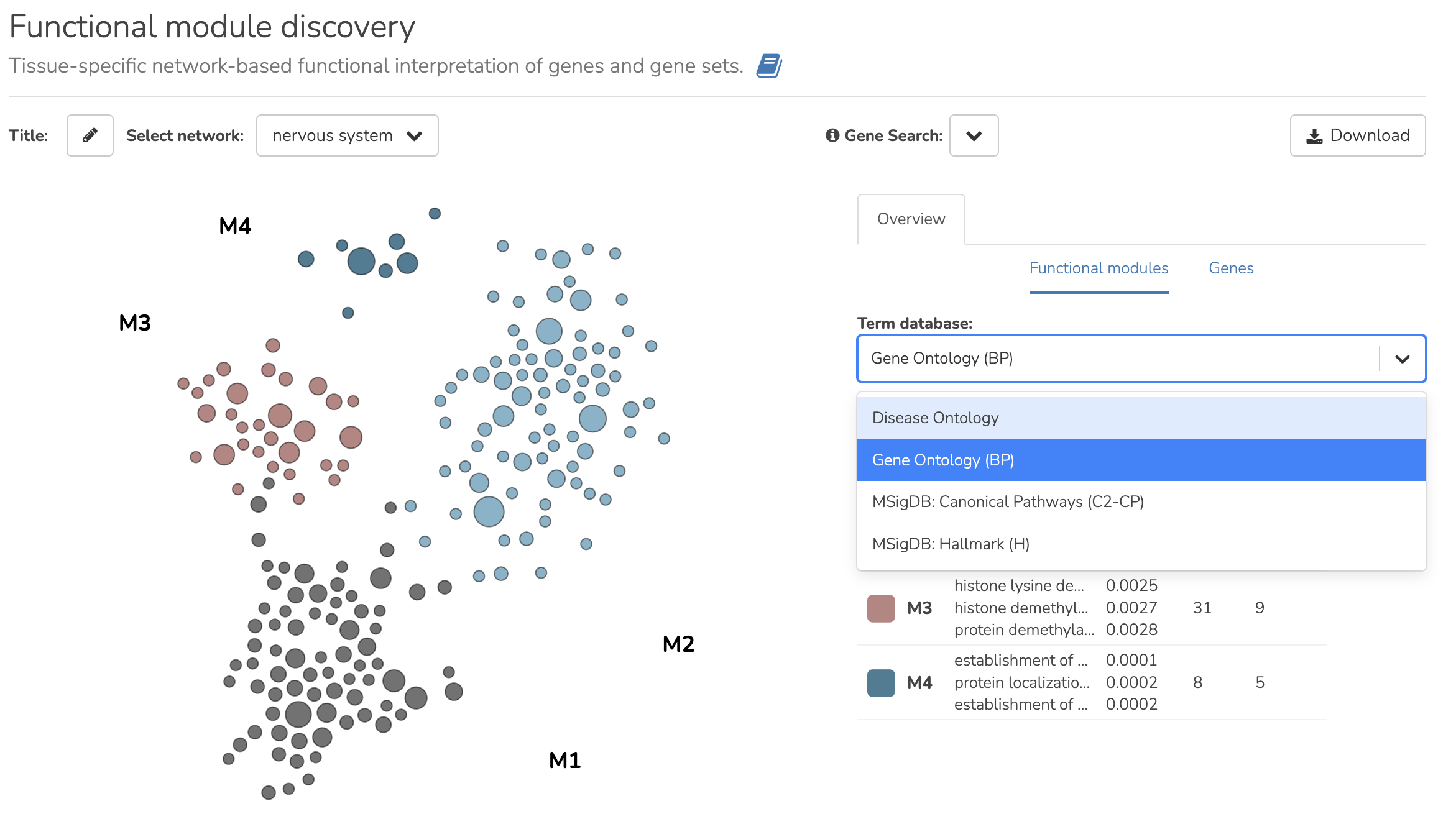
Expanded Enrichment Databases:
We have enhanced the Functional Module Detection tool by adding support for multiple databases. Previously, only Gene Ontology Biological Process (GO BP) terms were available. Now, you can explore enrichment analysis of the detected modules from the following four databases:
- Gene Ontology Biological Process (GO BP): Learn more about GO BP
- Disease Ontology: Learn more about Disease Ontology
- MSigDB Hallmark (H): MSigDB Collections
- MSigDB Canonical Pathways (C2-CP): MSigDB Collections
July 2024
Interactive API Documentation:
We added interactivity to all endpoints with working examples in the API documentation. This allows users to experiment with the API directly from the documentation page.
Check out the new API documentation here.
Functional Module Detection:
Functional Module Detection results may now be optionally named with a title.
The title will show in the browser as well as in the file name of the download files. There is a new breadcrumb navigation across all of HumanBase for better context while browsing the site.